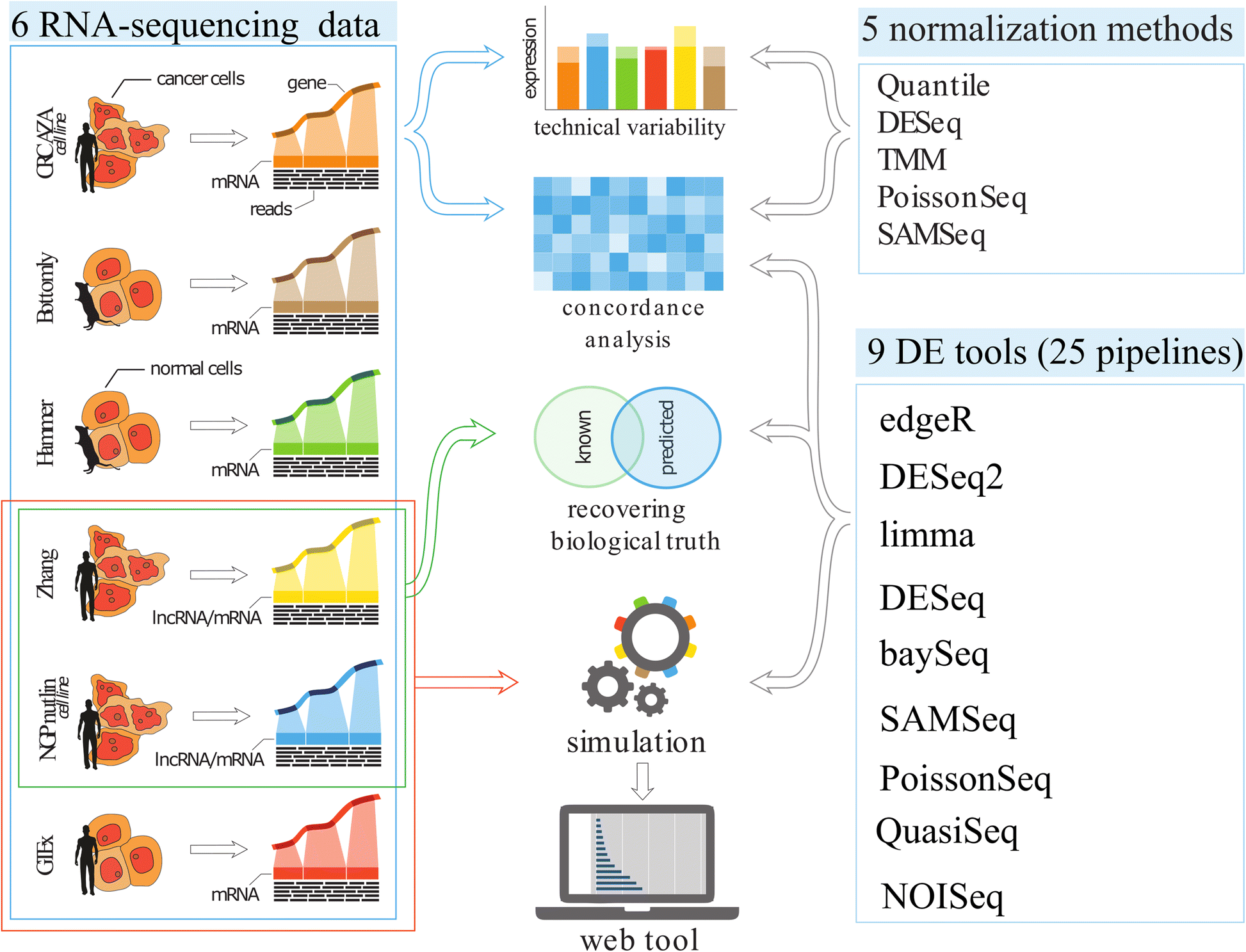
Differential gene expression analysis tools exhibit substandard performance for long non-coding RNA-sequencing data | Genome Biology | Full Text
A comparative study of RNA-Seq and microarray data analysis on the two examples of rectal-cancer patients and Burkitt Lymphoma cells | PLOS ONE
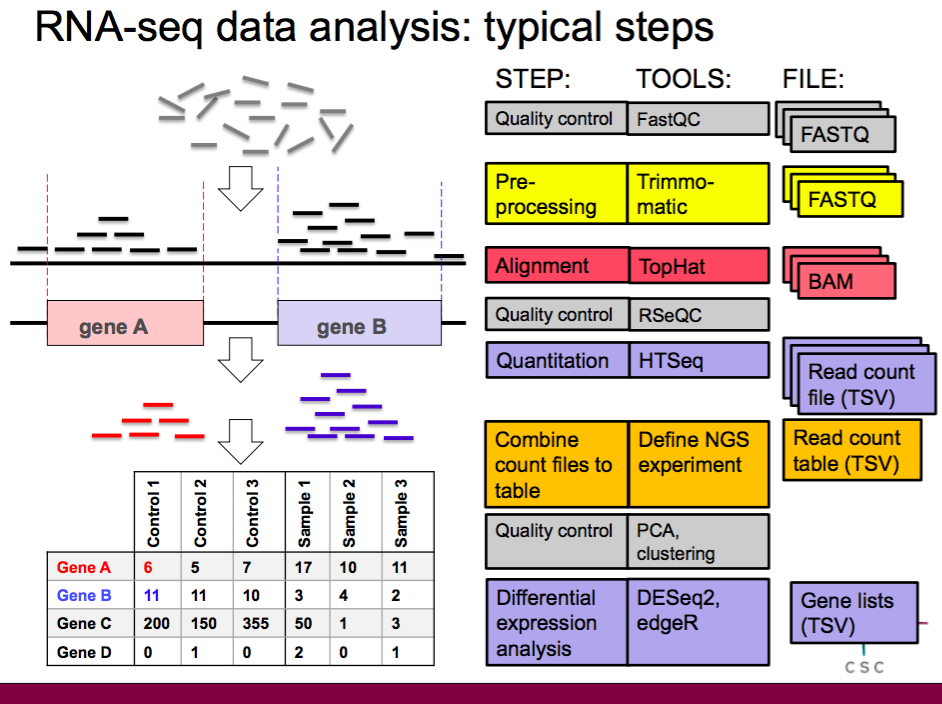
Summarizing RNA-seq alignment tools. (A) RNA-seq alignment: typical... | Download Scientific Diagram

Differential gene expression analysis tools exhibit substandard performance for long non-coding RNA–sequencing data | bioRxiv
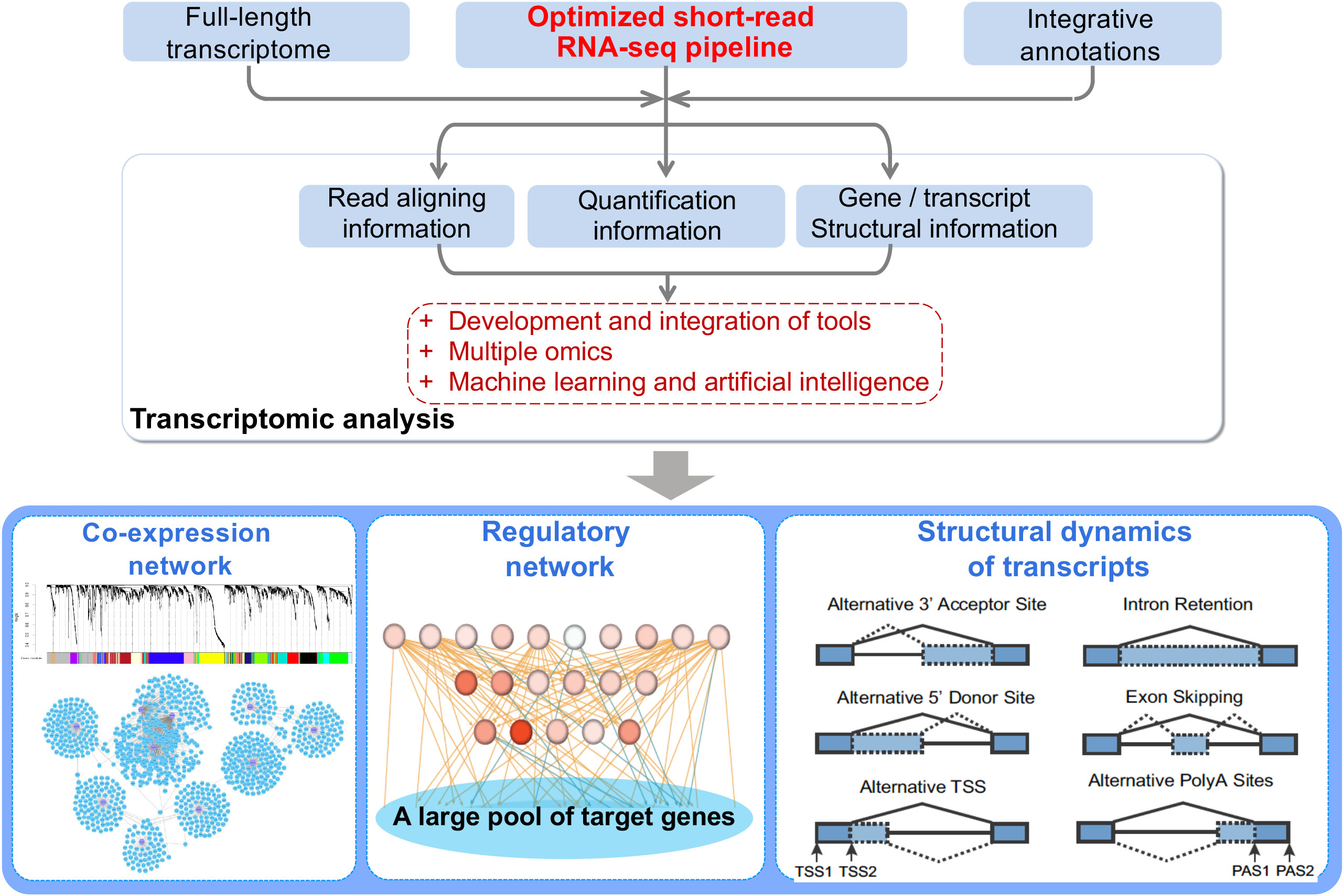
Frontiers | Unleashing the power within short-read RNA-seq for plant research: Beyond differential expression analysis and toward regulomics

cellxgene VIP – a web-based tool for the exploration and visualization of scRNA-seq data | RNA-Seq Blog

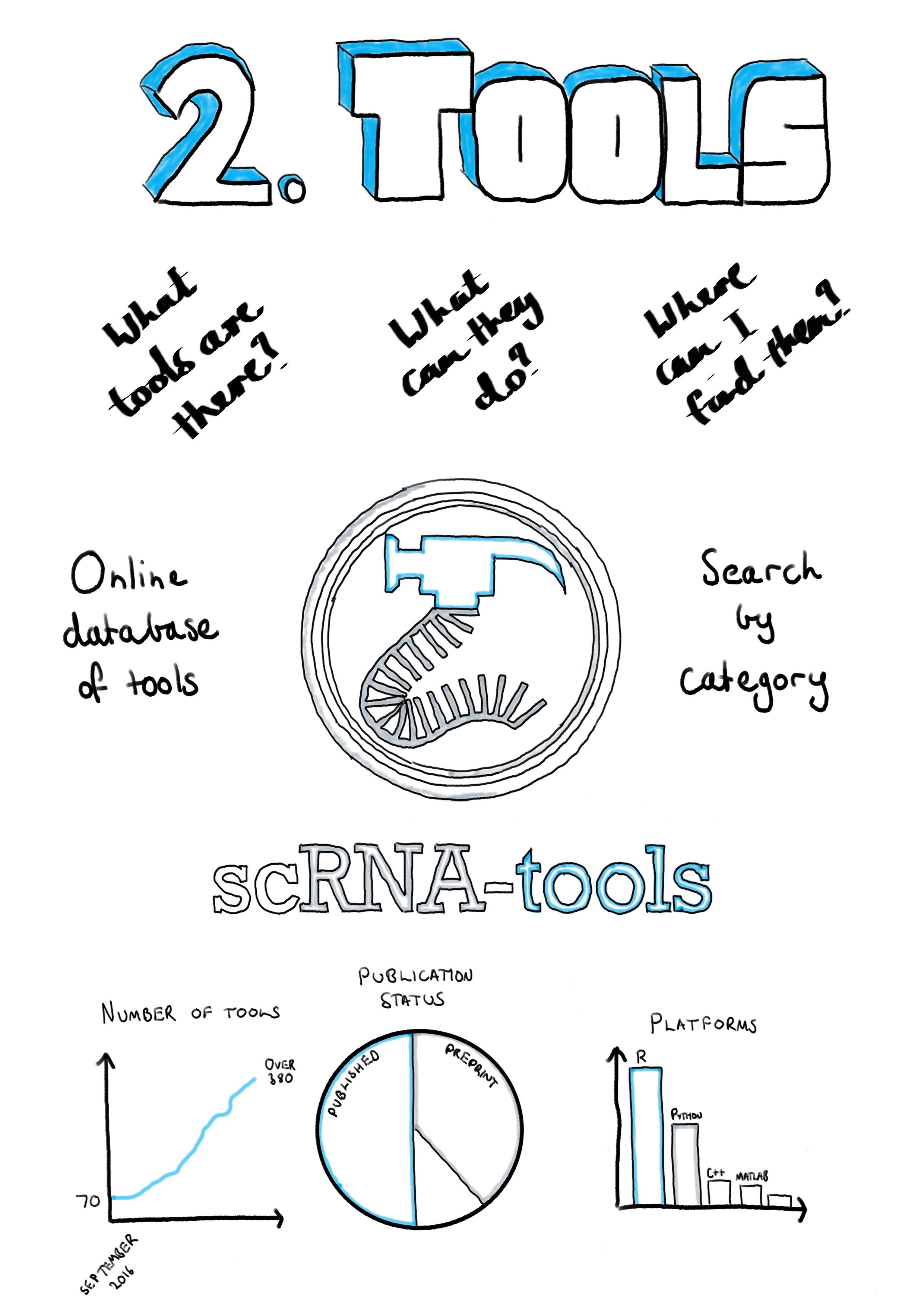
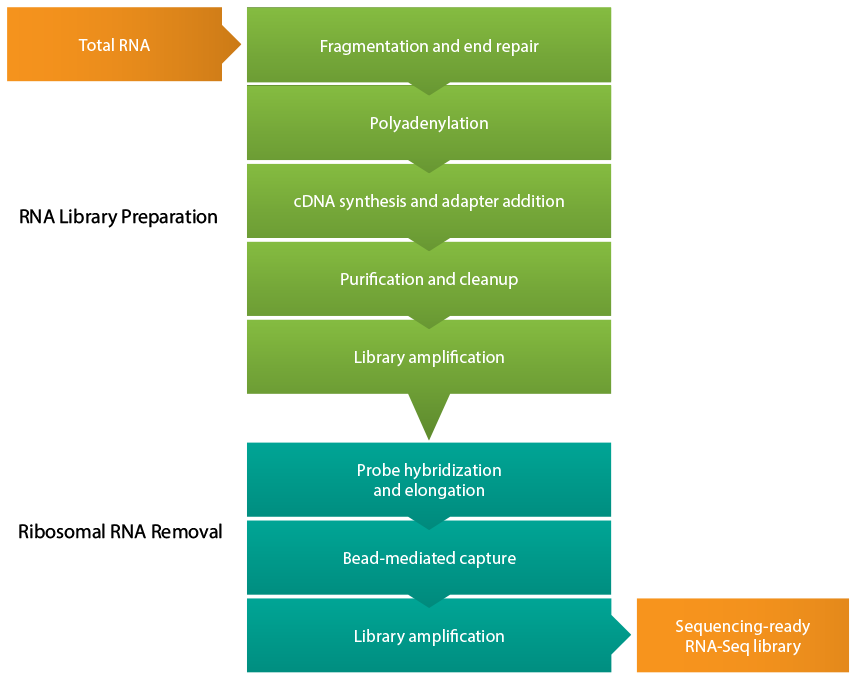
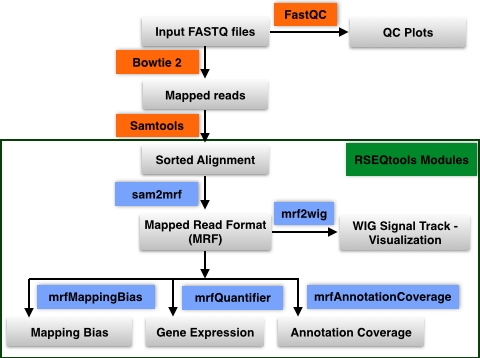
Learn more about available bioinformatics tools and pipelines to analyze RNA sequencing data | RNA-Seq Blog

Processing single-cell RNA-seq data for dimension reduction-based analyses using open-source tools - ScienceDirect
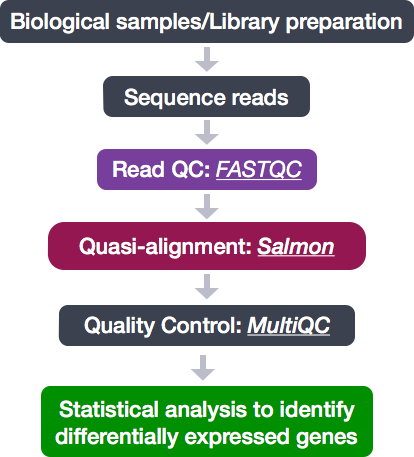



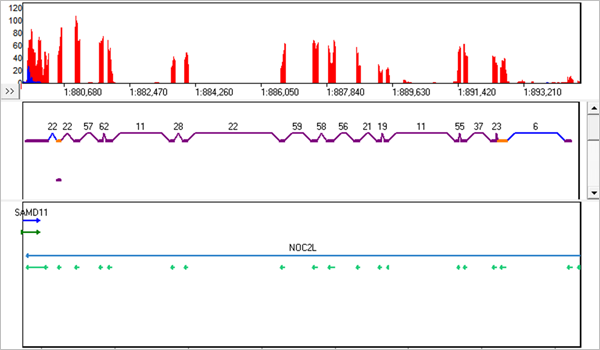





![PDF] A comprehensive overview of computational tools for RNA-seq analysis | Semantic Scholar PDF] A comprehensive overview of computational tools for RNA-seq analysis | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/76ec60bf27d5c242d8750674c44295b8bf56f413/44-Figure5-1.png)